Fig. 1
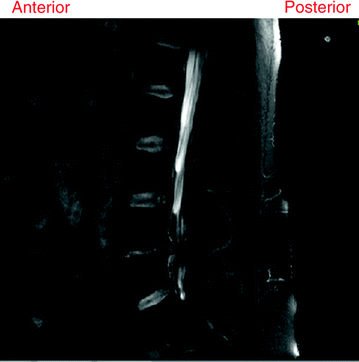
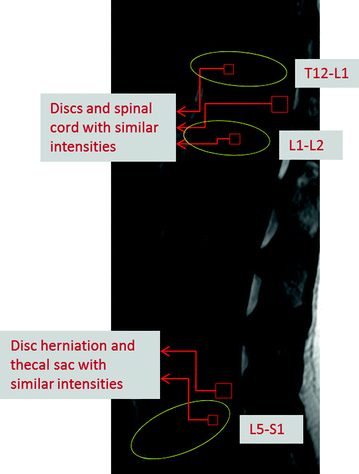
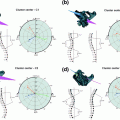
Localization and labeling of a sagittal lumbar T2-weighted MRI. Lumbar area is the second area to the last of the vertebral column. It is the main part of the vertebral column that is responsible for bearing the major body weight. The lowest lumbar vertebra is L5 and the highest is L1. Inter-vertebral discs are labeled based on the enclosing vertebrae [5]
The essential structures within the vertebral column that have been attracting researchers for localization, labeling, and segmentation are the intervertebral discs, the vertebrae and the Dural Sac.
The lumbar vertebrae are the five vertebrae between the rib cage and the pelvis which are designated as L1–L5, starting at the top. The intervertebral discs are fibrocartilaginous cushions which are named upon the vertebral bodies that sandwich a particular disc, e.g., the disc in between L1 and L2 is named L1-L2. In clinical practice, the radiologist reports the diagnosis at each disc level and at each vertebra level. Hence, the first requirement of any lumbar Computer Aided Diagnosis (CAD) system is to localize and label the lumbar discs and vertebrae as shown in Fig. 1 [5]. Specifically, localization refers to providing centroids or bounding boxes for each of the lumbar discs, and labeling refers to identifying each localized disc as one of the six lumbar discs (T12-L1, L1-L2, L2-L3, L3-L4, L4-L5, L5-S1). While some researchers have discussed methods to provide a point within each lumbar disc [5, 76], there are also methods [32] that provide a bounding box around every visible disc in clinical lumbar MRIs as illustrated in Fig. 2.
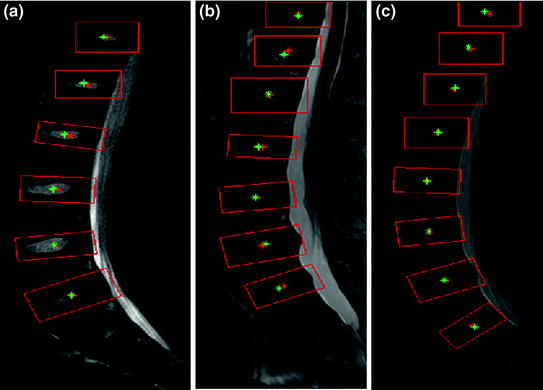

Fig. 2
This figure illustrates the results of an automatic lumbar disc localization method [32] which detects all the visible discs in a lumbar MRI. In case more than six discs are detected, the lower most six discs are identified as the lumbar discs. The red boxes are the bounding boxes provided for each of the visible discs in the clinical MRI. The red stars show the automatic disc centers, while the green stars show the true centers
Another important tissue structure in the lumbar spine is the Dural Sac. It is the membranous sac that encases the spinal cord within the bony structure of the vertebral column. The human spinal cord extends from the foramen magnum and continues through to the conus medullaris near the second lumbar vertebra, terminating in a fibrous extension known as the filum terminale. The Dural Sac usually ends at the vertebral level of the second sacral vertebra. Intensity inhomogeneity within the sac due to varying amounts of white and gray matter makes the segmentation of the Dural Sac and the spinal cord very challenging. Moreover, automatic segmentation in clinical MRIs is even harder due to variations in appearance and a lack of bright spinal fluid in cases with certain abnormalities such as stenosis.
After localization and labeling, comes the challenging task of tissue segmentation. Segmentation of discs is quite difficult due to extreme variability in shape, size and appearance of intervertebral discs in lumbar MRI. Moreover, discs with abnormalities can be very fuzzy and difficult to segment manually leading to significant inter-observer variability. At the same time, segmentation of intervertebral discs is a very important part of lumbar CAD systems in order to diagnose and quantify abnormalities such as herniation, desiccation and degeneration.
Requirements for CAD systems of the lumbar region are unique since we need to segment the Dural Sac and localize, label and segment the lumbar intervertebral discs before we can initiate the diagnosis. Figure 3 shows an illustration of automated segmentation [34] of the discs, vertebrae and the Dural Sac of a clinical MRI using two methods, the first using a probability map and HOG features, while the second method uses neighborhood label information as well, in a Gibbs Sampling approach.
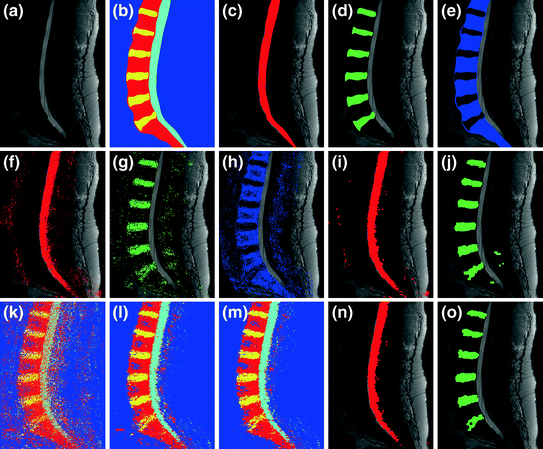

Fig. 3
Illustration of automated lumbar tissue segmentation [34]: a shows the original mid-sagittal MRI, b–e show the manual segmentation (ground truth), f–h show the label maps for the dural sac, disc and vertebra respectively using method 1 (probability map + HOG features), while i and j show the dural sac and disc segmentation after morphological post processing. k–m Show the label maps generated at the end of iteration number 1, 6 and 200 respectively using method 2 (probability map + HOG features + neighborhood labels via Gibbs sampling), while n and o show the dural sac and disc segmentation after morphological post processing
2 The Vertebral Column
This section is dedicated to present the anatomy of the vertebral column in general with focus on the lumbar area. It also provides the standardized nomenclature of the various abnormalities in the vertebral column as endorsed by the North American Spine Society (NASS), the American Society of Spine Radiology (ASSR), and the American Society of Neuroradiology (ASNR) [24].
The vertebral column, also known as the backbone or the spinal column, is typically made up of (33) individual bones called vertebra (plural: vertebrae) that interlocks with each other. These vertebrae are classified into five areas from top to bottom: Cervical (7), Thoracic (12), Lumbar (5), Sacral (5), and Coccyx (4). Among these (33) vertebrae, only the top (24) are movable due to which clinicians often state that the vertebral column consists only of (26) vertebrae counting the Sacral vertebrae as one and the Coccyx as one. In each of these four regions, the vertebrae have unique features that allows certain functionality [88].
There are five distinct regions in the vertebral column. The top most region is the Cervical region which consists of seven vertebrae anatomically named from top to bottom as C1 (also called Atlas), C2 (also called Axis), C3, C4, C5, C6, and C7. The main function of this region is to support the weight of the head [normally weighs about 10 pounds (4.5 kg)]. The cervical has the most range of motion due to the first two specialized vertebrae that connect to the skull.
The Thoracic region comes next and consists of twelve vertebrae named from top to bottom as T1, T2, T3, T4, T5, T6, T7, T8, T9, T10, T11, and T12. The major role of the thoracic spine is to protect the organs that lie in the chest by supporting the rib cage. The motion is limited due to the nature of the chest.
The Lumbar region comes next and has the largest vertebrae. This region is responsible for the whole flexibility of the back as well as bearing the weight of the body. Five vertebrae exist in this area that are named (from top to bottom) as L1, L2, L3, L4, and L5. The spinal cord stops, typically at the L1-L2 area where the nerves hang down inside the Pachymeninx (Dural Mater) which is the tough and inflexible outermost of the three layers of the meninges surrounding the brain and spinal cord. The Sacral and the Coccyx regions have less functionality and they are barely movable. The sacral one fused vertebra (named as S or S1) provides attachment for the Ilium (hip) bones and protects the pelvic organs while the coccyx region fused vertebra are not recognized for main functionality.
2.1 Lumbar Spine Region
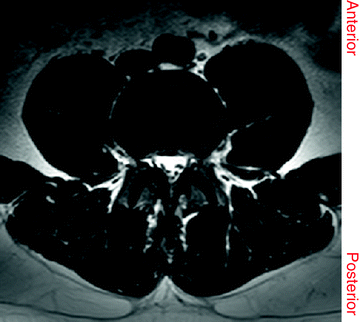
Since our focus is on the lumbar region, we present more details on the anatomy of the lumbar region as well as details of the MRI. There are two common anatomic terms that relate to the low back: anterior and posterior. Anterior refers to the front of the spine while posterior refers to back of the spine as shown in Figs. 4 and 5. The section of the spine that makes up the low back is called the lumbar spine. Lumbar spine includes these main structures: Intervertebral discs, vertebrae, and other structures.
2.1.1 Intervertebral Discs
Intervertebral discs are unique structures that absorb shocks between adjacent vertebrae. They act as the ligaments that connect the vertebrae together and the pivot point which allows the spine mobility by bending and rotating. They make about one fourth of the spinal column length [88].
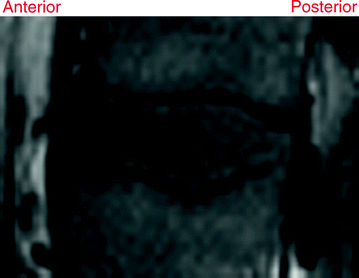
An intervertebral disc is composed of two parts: an outer strong ring called Annulus Fibrosus and a soft gel-like inner called Nucleus Pulposus. The nucleus pulposus consists of 80–85 % water in normal cases. By aging the disc dehydrates limiting its ability to absorb shocks. The outer rings gets weaker as well and start having tears that causes various abnormalities. The bottom up view is the anterior while top down is the posterior direction. Figures 6 and 7 show T1- and T2-weighted MRI for the same lumbar disc from our dataset, respectively.
2.1.2 Vertebrae
A vertebra (plural: vertebrae) is a bone with specific structure that supports and protects the spinal cord. A typical vertebra consists of two segments: Anterior (front) and posterior (back). The anterior part of the vertebra is the body while the posterior, which is known also as the vertebral (neural) arch, includes: the vertebral foramen, a pair of pedicles and a pair of laminae, and supports seven processes [four articular, two transverse, and one spinous (aka neural spine)].
2.1.3 Other Structures
In addition to the main structures within the vertebral column, there are few other structures including.
Nerves: The spinal cord hangs inside a bony ring through the vertebral column that is made up of millions of nerve fibers. The spinal cord extends down to the L2 vertebra. Below L2, a bundle of nerves named as Cauda Equina hangs down in what is known as the Thecal Sac. Two large nerves branch off the spinal cord, one from each side passing through the neural foramina of each vertebra. These spinal nerves group together to form the main nerves that go to the organs and limbs. The nerves of the lumbar spine (Cauda Equina) go to the pelvic organs and lower limbs [88].
Connective tissues: They are the fibrous connections that hold the cells of the body together. The ligaments are strong connective tissues that attach bones together. There are many long ligaments that connect on the front and back sections of the vertebrae. The anterior longitudinal ligament runs lengthwise down the front of the vertebral bodies. Two other ligaments run full-length within the spinal canal. The posterior longitudinal ligament attaches on the back of the vertebral bodies. The Ligamentum Flavum is a long elastic band that connects to the front surface of the lamina bones (just behind the spinal cord). Thick ligaments also connect the bones of the lumbar spine to the sacrum (the bone below L5) and pelvis.
Muscles in the lower back are arranged in layers. The superficial layer, the closest to the skin, is covered by a thick tissue called Fascia. The middle layer, called the Erector Spinae, has strap-shaped muscles that run up and down over the lower ribs, chest, and low back. They join in the lumbar spine to form a thick tendon that binds the bones of the low back, pelvis, and sacrum. The deepest layer of muscles attaches along the back surface of the spine bones, connecting the low back, pelvis, and sacrum. These deepest muscles coordinate their actions with the muscles of the abdomen to help hold the spine steady during activity [95].
Spinal segments is a notion that includes two vertebrae separated by an intervertebral disc, the nerves that leave the spinal column at each vertebra, and the small facet joints that link each level of the spinal column. The intervertebral disc separates the two vertebral bodies of the spinal segment. The disc normally works like a shock absorber. It protects the spine against the daily pull of gravity. It also protects the spine during heavy activities that put strong force on the spine, such as jumping, running, and lifting. The spinal segment is connected by two facet joints described earlier. When the facet joints of the lumbar spine move together, they bend and turn the low back [95].
3 Popular Lumbar Imaging Modalities
Most image based research literature focuses on X-ray radiography, Dual-energy X-ray Absorptiometry (DEXA or DXA), CT, and MRI. X-ray radiography and DEXA are cheaper and widely popular modalities as an initial diagnostic tool. Hence, the availability of the data provided researchers with great opportunities to investigate labeling, localization, and even diagnosis problems.
On the other hand, MRI (Fig. 8) and CT (Fig. 9) are more expensive and less available for researchers. Hence, fewer researchers obtained access to such data and were able to investigate localization, segmentation, and diagnosis problems on the various anatomical structures. Few efforts utilized other modalities such as ultrasound, especially, for fetal spine detection and abnormality detection.
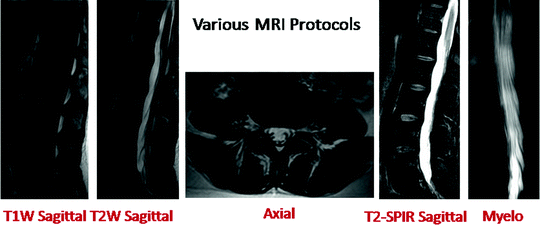
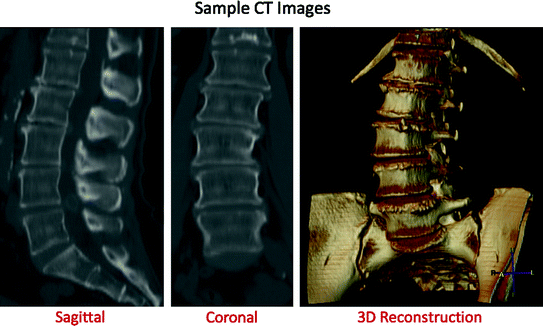

Fig. 8
This figure illustrates a sample image from each of the five popular clinical MRI protocols—T1 weighted sagittal, T2 weighted sagittal, T2 weighted axial, T2-SPIR sagittal and Myelo [6]

Fig. 9
This figure shows samples of a lumbar CT scan. The first two images show sagittal and coronal slices from a CT volume, while the last image shows a 3D reconstruction [2]
Both X-ray and DEXA (aka DXA) radiography consist only from one 2D slice that shows the area of interest. On the other hand, CT scans show a full 3D volume for the area of interest. Clinical CT spans the whole area in slice-by-slice fashion that can be directly used to produce a full 3D volume. Usually CT consists of a set of axial slices with specific thickness depending on the available technology.
Moreover, clinical MRI consists of few protocols that vary depending on the available technology. The current standard in MRI in North America, for low back, is the 3 T MRI. Most of the current MRI radiology centers produce: (1) T1-Weighted sagittal (T1 W-sagittal), (2) T2-Weighted sagittal (T2 W-sagittal), (3) T2-Weighted axial (T2 W-axial) for a set of selected discs, (4) T2-Weighted axial (T2 W-axial) and (5) Myelo MR images (Fig. 8). While the sagittal views span the side-to-side dimension of the body, the axial views are acquisitions of each intervertebral disc within the area of interest. Each disc has a set of axial slices that are aligned with the dimension of the major axis of the disc. MRI acquisition technician spends a manual effort for planning the acquisition to make sure that each disc volume is acquired correctly and that all acquired protocols are manually co-registered. The patient is not allowed to move during the whole acquisition period.
Many clinical MRI protocols exist that have trade-offs in diagnosis of various backbone abnormalities. The technician has four main parameters to tune before MRI acquisition that control the appearance (intensity) of the resulting image: (1) proton density, (2) longitudinal relaxation time (T1), (3) transverse relaxation time (T2), and (4) the flow. The proton density refers to the concentration of protons in the tissue in the form of water and macromolecules (proteins, fat, etc.). Both T1 and T2 relaxation times define the way that the protons revert back to their resting states after the initial RF pulse. The most common effect of the flow is the loss of signal from rapidly flowing arterial blood.
Two common pulse sequences for MR imaging are widely used: T1- and T2-weighted spin-echo sequences. The T1-weighted sequence uses a short Repetition Time (TR) and a short TE (Echo Time) (TR ≤ 1,000 ms, TE ≤ 30 ms). The T2-weighted sequence uses a long TR and long TE (TR ≥ 2,000 ms, TE ≥ 80 ms). Moreover, two major techniques are used for suppression of fat signals in MRI: Short Tau Inversion Recovery (STIR) and selective partial inversion recovery (SPIR). The STIR sequences suppress fat signal by using an initial 180° radiofrequency pulse to invert the longitudinal magnetization. Image acquisitions are then performed with the inversion time equivalent to the known null point for fat (approximately 0.69 × T1) [92]. SPIR is a more recent fat-suppression technique that is based on the use of frequency-specific pulse sequences [93]. Only the fat magnetization pulse is inverted leaving water resonances as is. This technique is useful for suppressing any tissue-specific pulse given the known-frequency of that tissue. However, the SPIR technique is extremely sensitive to the magnetic field inhomogeneity. SPIR is used with both T1- and T2-weighted MRI [44].
Furthermore, another important MRI sequence generator that is related to common current clinical MRI is called MR Myelography [57] (Myelo is a new Latin word, from Greek muelos, which means spinal cord). In this method, the background signal is suppressed by using heavily T2-weighted fast spin-echo pulse sequences and obliterating fat signal by pre-saturation. The resulting slices are then projected into a composite image using a standard maximum intensity projection (MIP) algorithm [57].
It is worth mentioning that inter-observer variability exist in lumbar diagnosis similar to many diagnosis tasks from various imaging modalities including X-ray radiographs, MRI, CT, Single-Photon Emission Computed Tomography (SPECT), and High Resolution (HR). However, MRI shows high inter-observer reliability compared to plain radiographs in lumbar area diagnosis (e.g., [62]). Mulconrey et al. [70] showed that abnormality detection for degenerative disc and Spondylolisthesis with MRI has κ = 0.773 and κ = 0.728, respectively, which is considered high in showing inter-observer reliability where this reliability is considered perfect when 0.8 ≤ κ ≤ 1.
4 Challenges
Automatic detection of abnormalities from MRI or CT scans has been studied by researchers for quite some time. The challenges are manifold—ranging from variations in scanner specifications, parameter settings, modalities, differences in body structure and composition and last but not the least the task of segmentation which is a big challenge in computer vision.
In general, the segmentation of CT and MRI scans is difficult due to three main reasons.
(1)
Partial Volume Effect: It is a scenario where multiple tissues contribute to pixels and blurs intensity across boundaries as illustrated in Fig. 10.
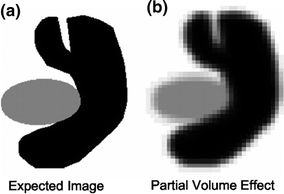

Fig. 10
This figure shows an illustration of partial volume effect in an imaginary scan consisting of two different kinds of tissues. While the first image shows the expected image the second one shows the actual image with fuzzy boundaries due to partial volume effect [32]
(2)
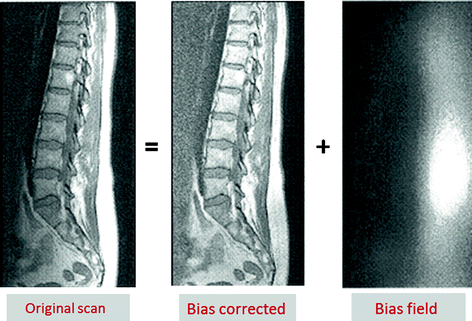
Intensity Inhomogeneity: It is defined as non-anatomic intensity variations of the same tissue over the image, and may be caused by the imaging instrumentation (RF non-uniformity, static field inhomogeneity) or due to patient movement as seen in Fig. 11.
All these factors contribute to the fact that segmentation of a lumbar MRI is a very challenging task.
Real world clinical MRIs are even more challenging since patients very often suffer from one or more lumbar abnormalities such as vertebral fractures, Spondylolysis, Spondylolisthesis, Scoliosis, intervertebral disc abnormalities (degeneration, desiccation, herniation, bulge and annular tears) and spinal Stenosis. In addition, there is lumbar variability due to patient age, height and structure leading to diverse images. Figure 13 shows a sample set of clinical lumbar MRIs showing some of the variability [5].
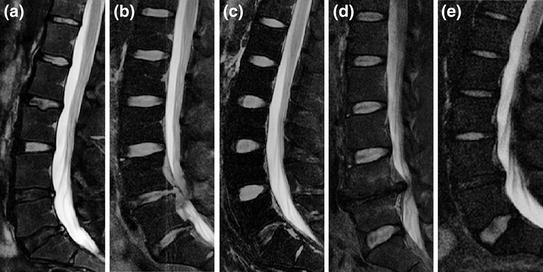

Fig. 13
Variability in disc appearances, shapes, locations, and sizes in different abnormal cases. a Shows variability in appearance of discs. The lower two discs (L4-L5 and L5-S1) have less intensity levels due to abnormalities (Herniation, Stenosis and Desiccation). b Shows variability in shape of discs with close intensity levels due to abnormalities in the lower two discs (Herniation and Stenosis). c Shows clear difference at the lowest disc level (L5-S1) as well as the difference in bending of the lumbar vertebral column which results in variability in location. d Shows variability in location of discs from other figures, sizes of discs, and the missing disc at L4-L5 disc. e Shows variability in disc sizes between the upper four discs and the lowest disc L5-S1. Ages of these patients are 35, 36, 29, 47, and 27, respectively from a to e. All images have been edited by cropping and contrast enhancement for better visualization
5 Advances in Localization, Labeling, and Segmentation
There are three main steps for the proper diagnosis in medical imaging: (1) Localization and labeling of anatomic structures, (2) Segmentation and (3) Diagnosis and quantification of abnormalities. There has been an extensive amount of work done in the area of vertebral body localization and segmentation from X-ray radiographs and CT scans in the past two decades. On the other hand, localization of soft tissues in MRI and diagnosis of disc abnormalities is comparatively more recent and has been of central focus for low back research in the last decade.
In this section, we review in detail, the current literature in the context of spinal tissue localization, segmentation and abnormality diagnosis. We classify the literature based on the medical imaging modality.
5.1 X-ray Radiography
Since X-ray radiographs use ionizing radiation and show better detailing of the bony tissues, there has been plenty of research in the direction of vertebrae segmentation from X-ray scans. Moreover, the availability of X-ray radiographs data helped boost the related amount of research.
5.1.1 Vertebrae
More than two decades ago, Hedlund and Gallagher [39] performed vertebral morphometry on lateral thoracic and lumbar X-ray radiographs of 153 women with a preliminary diagnosis of Spinal Osteoporosis. Measurements included anterior and posterior vertebral height, width, area, wedge angle, percent reduction of Anterior to Posterior Height (PRH) and Percent Difference in Anterior Height between adjoining vertebrae (PDAH). They showed that among individuals with mild Osteoporosis (0–2 fractures) PDAH identified 86 % of the fractures and 95 % of the individuals with fractures.
Manual selection of anatomical points for vertebral abnormality diagnosis is time consuming, imprecise and subjective. To obtain a more objective and accurate description of the vertebral body shape, semi-automatic methods were proposed that were based on statistical models of vertebral bodies in the sagittal view. Very early on, in 1993, a computerized quantifying technique for vertebral morphometry on lateral radiographs of the spine was proposed by Nicholson et al. [73]. Although fracture detection was improved by expanding the description of the vertebral body shape from six points to a contour, the amount of traumatic spinal injury or latent vertebral fracture was often underestimated. The main reason for the wrong diagnosis originated from the limited measurement possibility caused by the lack of depth perception in X-ray radiographs. Later in 1997, Smyth et al. [87] described how Active Shape Models (ASM) could be used to locate both normal and fractured vertebrae from Dual energy X-ray Absoptiometry (DXA) images of the spine. However, three initialization points have to be manually selected. To overcome the lack of depth perception in X-ray images, Benameur et al. [8] performed projection of a three-dimensional (3D) statistical shape model of a vertebra to a pair of orthogonal 2D X-ray radiographs. They validated this method on 57 scoliotic vertebrae images. However, the proposed segmentation was highly dependent on model initialization.
Various efforts that target the diagnosis of certain vertebra conditions involved localization and segmentation. In 2000, Long and Thoma [60] investigated the segmentation of C2 and C3 vertebrae from the cervical area using an ASM as a first step for building an image based retrieval system for a dataset consisting of 7,000 lumbar X-ray radiographs and 10,000 cervical spine X-ray radiographs. They built the Web-based Medical Information Retrieval System (WebMIRS) based on the National Health and Nutrition Examination Surveys (NHANES). Later, Cherukuri et al. [16] proposed image processing techniques for computing size-invariant, convex hull-based features to highlight anterior Osteophytes. Feature evaluation of 714 lumbar spine vertebrae using a multi-layer perceptron yielded normal and abnormal average correct discrimination of 90.5 and 86.6 %, respectively. Despite that the main purpose of this work is diagnosis, a great portion of this effort was to identify the vertebra.
Another work was presented by de Bruijne and Nielsen [11] who used Shape Particle Filter [23] and k-nearest neighbor pixel-level classification for a semi-automatic segmentation of the lumbar vertebrae. Around the same time, Kaminsky et al. [47] presented a standardized protocol that combined newly developed interactive tools (rotation transformation, warped dissection plane) with standard segmentation tools to provide both a fast and accurate 3D spine segmentation procedure. Howe et al. [42] proposed a multi-level segmentation technique for vertebrae from cervical and lumbar X-ray radiographs using an ASM and a generalized Hough transform. Their validation is based on a leave-one-out test with an error of 2 mm for 57 % of the cervical cases and less than 4 mm for 68 % of the lumbar cases. However, their method needs manual intervention for initialization.
Later, Crimi et al. [22] presented a Bayesian approach and used prior information to estimate the covariance matrix from a small number of samples in a high dimensional shape to segment the vertebrae from X-ray radiographs. Moreover, Zewail et al. [103] segmented the vertebrae from X-ray radiographs using a contourlet-based salient point matching and a localized multi-scale shape prior. They tested their work on 100 X-ray radiographs and obtained an average segmentation error of 1.2 mm. Later, Crimi et al. [22] presented a Bayesian approach and used prior information to estimate the covariance matrix from a small number of samples in a high dimensional shape to segment the vertebrae from X-ray radiographs. Moreover, Zewail et al. [103] segmented the vertebrae from X-ray radiographs using a Contourlet-based salient point matching and a localized multi-scale shape prior. They tested their work on 100 X-ray radiographs and obtained an average segmentation error of 1.2 mm.
More recently, Lecron et al. [59] presented a fully automated vertebrae detection. They used an edge polygonal approximation to detect vertebral edges and a SIFT descriptor to train an SVM-model. They achieved a corner detection rate of 90.4 % and a vertebra detection rate from 81.6 to 86.5 % on 250 cervical cases.
5.1.2 Intervertebral Discs and Dural Sac
Despite that X-ray radiographs are mainly used for bone visualization, few efforts have been performed in the analysis of the intervertebral discs and the spinal canal. Chamarthy et al. [14] introduced image analysis techniques, including scale-invariant, distance transform-based features to characterize the disc space narrowing (with four grades zero to three) in cervical vertebrae. For a data set of 294 vertebrae X-ray images, experimental results yielded average correct grade assignment of greater than 82.10 %. However, the testing images had to be manually labeled with the boundary points.
Later on, Koompairojn et al. [56] described a fully automatic Spinal Stenosis diagnosis system via vertebral morphometry [73], using an Active Appearance Model (AAM) for segmentation and a Bayesian framework for classification. Experimental results on 86 lumbar spine X-ray images from the NHANES II database showed accuracy ranging from 75 to 80 %. Moreover, Stanley et al. [89] investigated new size-invariant features (claw and traction) for the detection of anterior Osteophytes for efficient Content Based Image Retrieval (CBIR). Using a K-means clustering and nearest neighbor classification approach, average correct classification rates of 85.80, 86.04 and 84.44 % were obtained for claw, traction and anterior Osteophytes, respectively, on 390 cervical vertebrae.
5.2 Computed Tomography (CT)
CT imaging technique has become indispensable for diagnosis of spine abnormalities by providing a detailed 3D representation of the anatomy. Compared to X-ray radiography and MR imaging, CT proved to have higher sensitivity and specificity in the visualization of the bone structures.
5.2.1 Vertebrae
A number of automatic and semi-automatic methods for segmentation of vertebrae and vertebral structures in CT have been proposed [27, 50, 52, 64, 80, 84, 91, 96, 98, 102] over the last decade. On one hand some researchers proposed techniques that segment each vertebra separately [50, 61, 64, 84, 98], which might lead to mis-segmentation due to absence of a clear boundary between vertebrae. To overcome this issue, some authors proposed techniques for simultaneous segmentation of all vertebrae [52, 80].
Early last decade, Hahn [36], proposed a fully automated approach to evaluate rotation of the cervical vertebrae in 3D using a multidimensional Powell minimization algorithm for spiral CT scans. Later, in 2004, the same research group [37] presented a method for determination of the planes separating the individual vertebrae of the spine from CT volumes using a Balloon based model. This model requires careful initialization similar to the 2D active contours (snakes) besides its high dependency on the edge detector.
Meanwhile, Ghebreab and Smeulders [27] presented a combination of Strings [26] and Necklaces [28] to model the spine in the lumbar area using both a priori knowledge about natural variation and anatomical saliency in the visual appearance of the spine. The Strings model focuses on learning the most relevant biological variation in the visual appearance of the spine as a whole, and Necklaces aims at exploiting inhomogeneities in multiple continuous shape and gray-level features of vertebrae. Thus they were able to use both a priori knowledge about natural variation and anatomical saliency in the visual appearance of the spine. However, they tested their method on only six CT cases and with minimal spinal and vertebral deformations. Furthermore, manual intervention is used for initialization of their model. On the other hand, Vrtovec et al. [96] detected the spine curve from CT using a polynomial model to provide a Curved Planar Reformation (CPR) of the 3D spinal column. They fit the spinal curve to a set of points extracted from a distance map that emphasized the vertebral bodies and tested the method on five cases, including one Scoliotic case achieving mean positional errors between 2 and 6 mm.
Furthermore, Yao et al. [102] presented a systematic algorithm for segmenting the spinal column from chest and/or abdominal CT scans, without labeling of vertebrae. Their method is based on thresholding, Watershed, and directed graph search besides modeling the vertebral bony tissue as a four-part model. They showed correct segmentation of 69 cases out of the 71 total cases. Meanwhile, Mastmeyer et al. [64] segmented the lumbar vertebral bodies in CT images by combining viscous deformable models with the geometrical shape of the vertebral body, starting from a point in the center of each vertebral body. Tan et al. [91] presented a level set-based segmentation algorithm for the vertebrae and validated their work on synthetic 3D vertebrae volumes. After parameter selection, they tested the algorithm on 50 vertebrae (from ten subjects), obtaining 90 % success rate. Later, Shen et al. [84] presented a segmentation technique for vertebrae from 3D CT scans using prior knowledge with a set of high level features to form a surface model. However, they did not perform labeling. They tested their model on 150 vertebrae with a comparative segmentation to two experts’ segmentation. In the same year, Klinder et al. [51] presented a two-scale framework for modeling and segmenting the spine from thoracic CT scans, achieving a segmentation accuracy of 1.0 mm in average for ten thoracic CT volumes. By applying statistical models of shape, gradient and appearance of spinal structures in 3D, the same research group [52] detected, identified and segmented the vertebrae in CT volumes. However, their identification algorithm is based on vertebrae Active Appearance Model for spatial registration and matching which is very computationally expensive (20–30 min per case). Their framework was tested on 64 CT images including pathologies like Scoliosis, Kyphosis and collapsed vertebrae. Later, Kim and Kim [50] automatically segmented the vertebrae by a region growing algorithm inside a volume limited by a 3D fence that was obtained from a deformable model. They obtain 80 % success on a 50 patient dataset. More recently, Ma and Lu [61] proposed a method for segmentation and identification of thoracic vertebrae in CT images by training an edge detector to bone structures via steerable gradient features and using a deformable surface model in a two-stage coarse-to-fine scheme. They achieve point-to-surface error 0.95 ± 0.91 mm on 40 volumes.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree