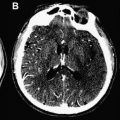
Although acute stroke imaging has made significant progress in the last few years, several improvements and validation steps are needed to make stroke-imaging techniques fully operational and appropriate in daily clinical practice. This review outlines the needs in the stroke-imaging field and describes a consortium that was founded to provide them.
The need for standardization and validation
Since the approval of intravenous tissue plasminogen activator (IV tPA) in 1996, the number of acute stroke patients receiving this agent has remained less than 5%, mainly because of delayed presentation to care. A recent study suggested that the time window for tPA administration could be extended from 3 to 4.5 hours in certain patients. Although this extension might slightly increase the rate of tPA administration, it does not change the fundamental limitation, which lies in the fact that currently the decision to initiate thrombolytic therapy depends on a generalized statistical rule regarding time to presentation rather than on an individualized pathophysiologic assessment of the ischemic penumbra, the target of tPA treatment. Several studies have suggested and validated the concept that the ischemic penumbra can be imaged and quantified, and an optimal therapeutic decision regarding thrombolytic agents can be based on such imaging, within an extended time window of up to 9 hours. Such an approach would not only increase the fraction of acute stroke patients amenable to thrombolytic treatment, possibly up to 40%, but also might allow more precise patient selection, thereby improving overall clinical outcomes.
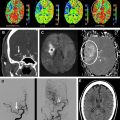
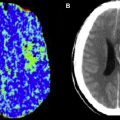
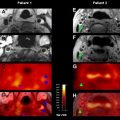
However, this concept of penumbral image-guided thrombolytic therapy has not yet been validated in a large phase 3 trial and as a result, has not become part of the clinical standard of care in tPA treatment decisions. One of the reasons for this delay in validation is the lack of standardization in penumbral imaging, which can result in different treatment decisions in the same patient when different penumbral imaging methods are applied to the same perfusion imaging data set. Current methods available for defining the penumbra vary for several reasons. The mathematical algorithms that calculate variables such as cerebral blood volume (CBV), cerebral blood flow (CBF), and mean transit time (MTT) differ. The variables that are used to define the infarct core and penumbra (eg, MTT vs CBF vs CBV in perfusion computed tomographic [CT] imaging) and their corresponding thresholds also differ ( Figs. 1–3 ).
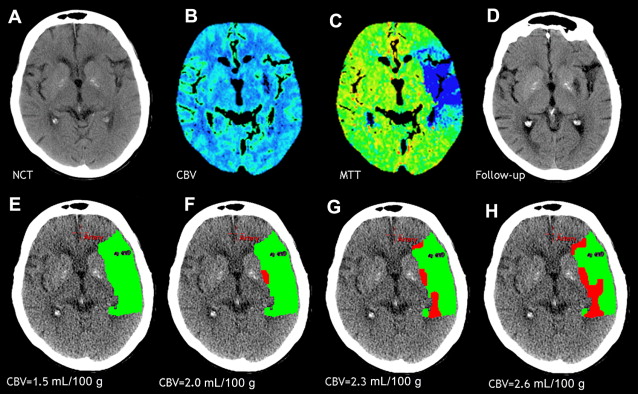
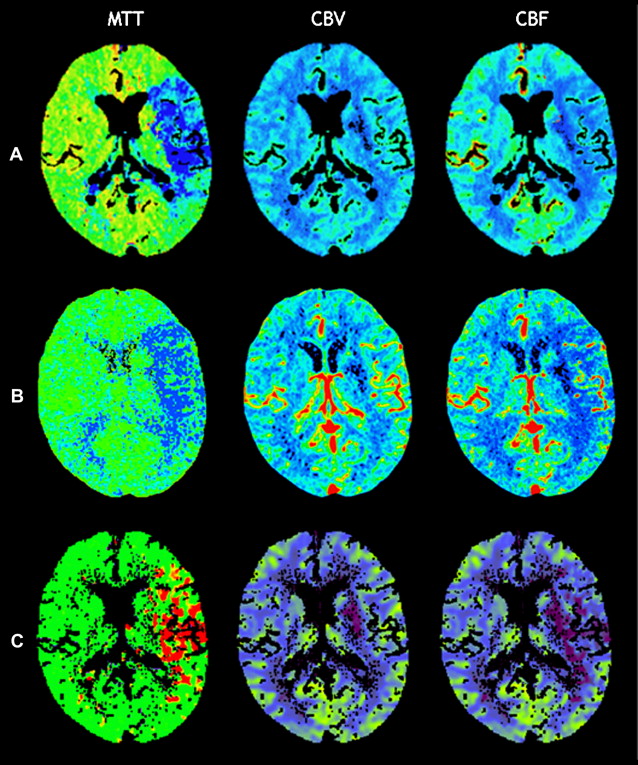
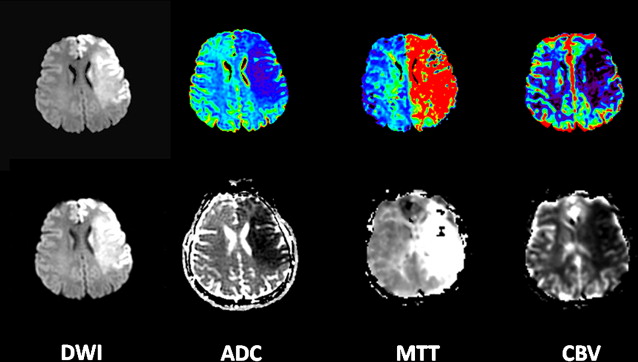
The differences between the imaging methods that identify the ischemic penumbra are not irreconcilable. Rather, they just need some refining to give concordant results, which is one of the goals pursued by the Stroke Imaging Repository (STIR) consortium.
The STIR
History
In September 2007, the National Institutes of Health, in conjunction with the American Society of Neuroradiology and the Neuroradiology Education and Research Foundation, sponsored a research symposium entitled “Advanced Neuroimaging for Acute Stroke Treatment.” This meeting brought together stroke neurologists, neuroradiologists, emergency physicians, and neuroimaging research scientists to discuss the role of advanced neuroimaging in acute stroke treatment. In particular, the goals of the meeting were to discuss unresolved issues regarding (1) the standardization of perfusion and penumbral imaging techniques, (2) the validation of the accuracy and clinical utility of imaging biomarkers of the ischemic penumbra, and (3) the validation of imaging biomarkers relevant to clinical outcome.
One of the recommendations was the creation of an international consortium of investigators, the STIR, to combine efforts and promote excellence in stroke care and stroke trial design and more specifically to overcome the issues faced by advanced stroke imaging, as mentioned earlier.
This central repository was designed to work in collaboration with 2 other collaborative groups, namely, Virtual International Stroke Trials Archive (VISTA) and MR Stroke. VISTA is an international collaborative venture that was established to facilitate the planning of randomized clinical trials and collects and gathers data sets from multiple institutions. The VISTA database provides the possibility to access a large volume of patient data on which original analyses can be performed, which would ultimately aid clinical trial design and development. MR Stroke is a collaborative effort between leading clinicians and clinical scientists around the world with a common interest in Magnetic Resonance Imaging in Stroke. MR Stroke is intended to facilitate the sharing and exchange of information and ideas, to provide access to a range of Web-based MR imaging tools and resources relevant to stroke, and to establish a forum for the exchange of information relevant to research using MR imaging in stroke.
Overall Goals
The overall purpose of the STIR is to create a repository of source MR images and CT images that could be used toward the objectives of standardization and validation of image acquisition, of image analysis, and of clinical research methods for image-based stroke research. More specifically, the consortium was originally established to collect and provide a rich resource of patient data sets to contribute meaningful information regarding the goals listed earlier: (1) the standardization of perfusion and penumbral imaging techniques, (2) the validation of the accuracy and clinical utility of imaging biomarkers of the ischemic penumbra, and (3) the validation of imaging biomarkers relevant to clinical outcomes.
Main Ongoing Tasks
To achieve these goals, the STIR consortium is focusing its initial efforts and resources on 3 main tasks.
Task 1: review of the literature
A systematic review of the stroke perfusion literature is needed to assess the different parameters and thresholds that have been used to define the infarct core and penumbra in acute stroke patients imaged with CT and/or MR perfusion imaging in prior studies or clinical trials. This systematic review will provide an approximation of the accuracy of each of these methods and the modifications required to optimize and standardize these methods.
Task 2: mathematical models and processing algorithms
This task involves establishing a list of the key postprocessing components typically implemented in perfusion software (motion correction, deconvolution, selection of the arterial input function, types of maps produced, and methods to calculate infarct core and penumbra), describing the types/values of each of these components in the different perfusion software packages (eg, type of deconvolution, delay sensitivity, number of arterial input functions), and finally, assessing the acceptable deviations for the different perfusion software components.
Ideally, the resultant standard perfusion software would fulfill the following premises: (1) the perfusion postprocessing would be completely automated and available either on the scanner console or on a remote workstation, (2) the perfusion software would generate an automated delineation of the infarct core and penumbra and produce an automated calculation of their volumes, (3) image processing and generation of the maps and volumes would take less than 5 minutes, and (4) processing would be robust, despite the use of slightly different image acquisition protocols and different degrees of image quality and noise.
Task 3: clinical validation of MR imaging and CT
The different perfusion software packages available, both academic and commercial ones, should be systematically compared and their results harmonized. In order to achieve this goal, a large data set of cases for each modality (MR imaging and CT) is collected. These cases are randomly divided into 2 groups. The first set of cases (training set) is analyzed separately from the second set (validation set). The training set is used to compare each software package to the same gold standard data set and to determine the required modifications specific to each software package. After this initial phase of refining, the software packages are “locked,” and the improved version is used to analyze the validation data set. The results of this analysis are collected and interpreted centrally, in an independent fashion, to validate the improved locked version of the different perfusion software packages and to show that all the software packages lead to concordant standardized results, appropriate to use in a stroke treatment trial.
Next Steps and Other Needs in the Field
Single institutions contributing to the central repository create a network that will apply standardized image acquisition protocols, allowing the constitution of a large homogenous data set. This image data set facilitates sample size calculations when planning a trial. In addition, studies to determine the optimal imaging modality and the optimal timing to perform imaging to predict clinical outcomes are made possible. If shorter follow-up periods are validated to assess outcome and if imaging is demonstrated as an accurate diagnostic method, the feasibility of stroke treatment clinical trials will be greatly enhanced and their cost will be decreased.
All these steps will facilitate and, hopefully, contribute to a successful phase 3 clinical trial of stroke thrombolytic therapy in an extended time window. This development will, in turn, allow a significantly increased number of acute stroke patients to benefit from thrombolytic therapy and will reduce the burden associated with stroke in the society.
Individual Projects
Analyses and/or research projects taking advantage of the STIR data set can be proposed by any STIR member through a written application sent to the Steering Committee.
Current individual projects or study proposals to the consortium that are related to imaging techniques, models, standardization, and validation include the follwing.
Tissue Infarction Risk Maps Applied to Multicenter Stroke Data (Project Leader: Ona Wu, PhD, Boston, MA)
This study proposes the application of a voxel-wise approach, to define the infarct core and the potentially salvageable tissue, to a large multicenter acute stroke series for model development and validation. Preliminary results based on single center data have shown that these algorithms can be used for tissue characterization. Traditionally, the presence of mismatch between diffusion-weighted MR imaging (DWI) and perfusion-weighted MR imaging (PWI) lesion volumes has been considered as the hallmark of potentially salvageable tissue. The use of the PWI-DWI mismatch, however, has been criticized as being suboptimal because of the lesion heterogeneity in both DWI and PWI that can possibly confound accurate assessment of tissue viability within regions of ischemic tissue. In comparison, voxel-wise methods, such as tissue risk maps, may provide a more sensitive and specific approach for identifying potentially salvageable tissue.
Improved Assessment of Infarct Size on Diffusion Magnetic Resonance Imaging; Use of the Graph-Cut Algorithm (Project Leader: Peter J. Yim, PhD, New Brunswick, NJ)
This study proposes the use of a novel approach to medical image segmentation, a graph-cut algorithm, for estimation of the infarct size by diffusion MR imaging to overcome the subtle but potential meaningful differences obtained from manual methods. The main hypotheses to be tested are (1) this method can reduce interobserver variability in the segmentation of cerebral infarct size obtained from diffusion-weighted MR images and (2) the infarct size estimated by this algorithm from DWI is predictive of final infarct size as assessed at follow-up imaging.
Predicting Final Extent of Ischemic Infarction Using Artificial Neural Network Analysis of Multi-parametric MR imaging in Patients With Stroke (Project Leader: Hassan Bagher-Ebadian, PhD, Detroit, MI)
In ischemic stroke, the final size of the ischemic lesion is the most important correlate of clinical functional outcome. This study proposes the application of an artificial neural network, to predict the final infarct volume, to a large data set of treated acute stroke patients. Preliminary results on stroke patients from a single center, who did not receive thrombolytic therapy have shown that this approach, using baseline MR images, produced a map that correlated well with the T2-weighted abnormality at 3 months. Because this model was trained to predict tissue fate in the absence of any treatment, its performance in treated patient populations remains to be assessed. The proposed study is intended to apply the trained artificial neural network to treated patients (preferably tPA-treated patients) and validate its results in a large patient population.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree