- A brief history of the discovery of DNA
- The general structure of DNA and RNA and their relationship to each other
- Relationship between genes and codons
- DNA replication and repair and some ramifications of faulty repair
- Differences between genome instability and epigenetics
Introduction
DNA is the blueprint for life. Blueprints contain all the necessary information needed for the construction of a building. The drawings include lengths of studs, square footage of each room, and location of the stairs. However, blueprints do not contain directions on how to combine which chemicals to grow a tree. There are also no instructions for manufacturing the laminate flooring located on the stairs. This is the main difference between a building blueprint and DNA. DNA contains all the required recipes for each component of the human body. The body cools itself using sweat glands to produce perspiration. Sweat gland construction utilizes specific genes for manufacturing the proteins used for plasma membranes. These genes, located in the DNA, supply the instructions for the complete construction of the cell and directions for how the cell performs its function. As previously discussed, the cell contains all the processes required for life and is considered the smallest unit of life.
In this chapter, the structure and function of DNA will be addressed in more detail. DNA structure begins with simple molecules and ends with a series of complex bonds. As for function, DNA carries the basic information of life, but transfer of that information undergoes the lengthy processes of transcription and translation.
History
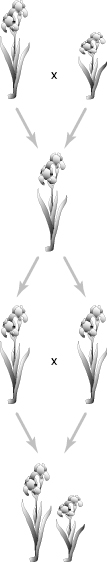
Although the study of genetics developed during the 20th century, its roots are based on studies from the late 1800s performed by Gregor Mendel. Mendel’s research, observation of pea plants in the monastery garden, was performed in relative obscurity. He studied the plants by following physical traits that were carried from one generation to the next. Eventually, Mendel began to interbreed pea plants with differing characteristics to see which would be passed to the next generation. With further observation and interbreeding, Mendel began to propose that each gene, or hereditary trait, was composed of two parts, known now as alleles. Mendel’s first breeding of pea plants crossed tall pea plants, growing 2 m in height, with short pea plants, growing only half a meter. The next generation of pea plants was tall, indicating two forms of alleles. The short trait was not seen. As the second generation of pea plants were interbred and grown, the result was a mixture of tall and short plants. Figure 6.1 shows Mendel’s pea plant experiment. These results confirmed Mendel’s theory of hereditary factors existing in two forms. In 1866, Mendel published his discoveries, but the article was not much noticed and he went on to do other things. Sixteen years after Mendel’s death in 1886, his paper was revisited. The study of genetics, as a science, was born and Mendel’s research technique was applied to many organisms.
Figure 6.1 A depiction of Mendel’s pea plant experiment. Tall and short plants were crossbred and all progeny were tall. However, crossbreeding the next generation resulted in both tall and short traits. Physical characteristics, such as height, can be easily tracked with minimal effort.
Although Mendel was able to demonstrate inheritance of physical characteristics, his studies did not pursue what was at the root of these traits. In the late 1800s, a German medical student, Johannes Friedrich Miescher, was interested in studying physiological chemistry using lymphocytes. However, he was encouraged by his mentor, Felix Hoppe-Seyler, to study leukocytes instead. Lymphocytes were difficult to obtain in sufficient quantities for study and leukocytes were readily available from puss. Miescher devised a system to isolate leukocytes, from a local hospital’s used bandages, without damaging the cells. A series of salt solutions washed the cells from the bandages and isolated the leukocytes. Once the cells were isolated, Miescher lysed the outer membrane and separated the nucleus for further research. When the nuclear membrane was lysed, a gelatinous material precipitated. Miescher called the precipitate nuclein, now known as DNA. As Miescher and his students studied the chemical structure of the nuclein, they were able to determine it contained large amounts of phosphorus and nitrogen but no sulfur. The results, being unlike anything seen before, caused Hoppe-Seyler to repeat all of Miescher’s experiments for confirmation. The results were published in Hoppe-Seyler’s journal in 1871.
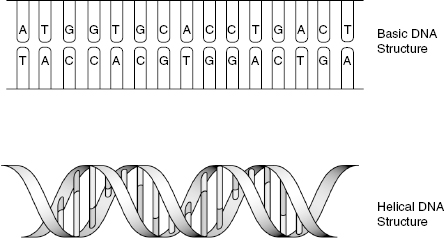
With the discoveries of inheritance and a nuclein made of phosphorus and nitrogen, the big question became “What is a gene?” In 1953, James Watson and Francis Crick tried to answer that question. Watson and Crick had studied DNA and knew nucleotides were connected together. These linkages were the product of chemical bonds between phosphate and sugar molecules located in the nucleotide itself. By linking the nucleotides together, a chain is formed and contains a particular sequence unique to that chain. The sequence differentiates each chain. Having this knowledge in hand, Watson and Crick proposed that DNA molecules consist of two chains of nucleotides, and these chains were held together by weak chemical bonds. In addition to proposing double-stranded DNA, Watson and Crick discovered that the two strands of DNA were wound around each other in a helical configuration. The structure of DNA, with and without helical arrangement, is shown in Fig. 6.2. Although the structure of DNA was determined, the idea of separate genes that encode traits was still being investigated.
Figure 6.2 Basic structure of DNA. The top picture demonstrates the association of nucleotides in a nonhelical fashion. The bottom picture shows the double-helix structure formed by DNA.
In the early 1900s, geneticists were working at identifying what genes were made of. After Watson and Crick discovered the structure of DNA, geneticists began to work on ways to determine the sequence of bases in DNA molecules. By obtaining the sequence of bases, or sequencing the DNA, all the information necessary to analyze the organism’s genes should be present. The collection of DNA molecules that is characteristic to an organism is referred to as its genome. Genome sequencing began with bacteria. Following this success, the Human Genome Project began in 1990 and was a worldwide effort to sequence the approximately 3 billion nucleotide pairs in human DNA. Upon completion of the Human Genome Project in 2003, the human gene number has been placed between 20,000 and 25,000 genes. These genes have been cataloged by location, structure, and potential function. Now, efforts have shifted to the discovery of how genes influence characteristics of the human being.
DNA Structure
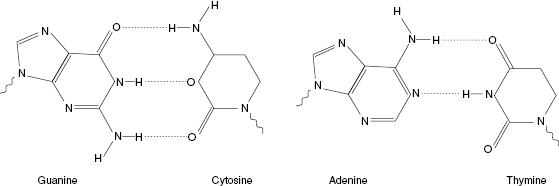
DNA, or deoxyribonucleic acid, is composed of a series of repeating units called nucleotides. Nucleotides consist of carbon, oxygen, hydrogen, nitrogen, and phosphorus. From these molecules, three basic elements are formed and combined to make a single nucleotide. These elements are nucleobase, carbohydrate, and phosphorus units. The nucleobase is made from a combination of nitrogen and carbon atoms that form either five- or six-member rings. Nucleobases are involved in the pairing of DNA polymers, known as complementary base pairing. In the DNA double helix, complementary base pairs occur between nucleobases of opposing DNA strands, which are bound together through hydrogen bonds. Hydrogen bonds are weak and can be broken and rejoined with relative ease. Complementary base pairing only occurs as indicated in Fig. 6.3. Adenine pairs uniquely with thymine (or uracil) and cytosine pairs with guanine. These specific interactions are critical for all functions of DNA. They help maintain the sequence of DNA throughout replication and allow reversible interactions to occur between the bases.
Figure 6.3 Complementary base pairing, indicating hydrogen bonds (dotted lines) between complementary nucleobase pairs. G-C base pairs always contain three hydrogen bonds, while A-T pairs contain two hydrogen bonds. This pairing is invariable throughout all DNA.
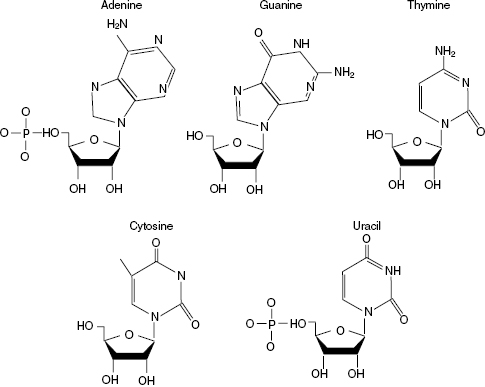
There are five nucleobases, and they are adenine (A), guanine (G), cytosine (C), thymine (T), and uracil (U). Three of the nucleobases are found in both DNA and RNA; however, DNA and RNA each have one unique base. Thymine is found solely in DNA and uracil in RNA. Each nucleobase is bound to a carbohydrate, or sugar, and at least one phosphate group. The carbohydrate is a pentose, or five-carbon, sugar. Deoxyribose is found in DNA while ribose is in RNA. The phosphate is simply a single phosphate atom surrounded by at least four oxygen atoms. With the addition of both a carbohydrate and phosphate, the nucleobase becomes known as a nucleotide. Nucleotides are the basic building blocks of DNA and RNA and will be the focus of further discussion. The basic structure of each nucleotide is shown in Fig. 6.4. Uracil, found only in RNA, contains a ribose sugar while adenine, guanine, cytosine, and thymine are shown with deoxyribose sugar.
Figure 6.4 Nucleotide structure. All nucleotides shown, except uracil, contain deoxyribose as the carbohydrate. A phosphate group is attached to the carbohydrate in both DNA and RNA polymers. Uracil, found only in RNA, replaces the deoxyribose with a ribose carbohydrate. Nucleotides utilized for RNA contain ribose in place of deoxyribose.
Nucleotides are held together through both ester and phosphodiester bonds. These bonds are much stronger than the hydrogen bonds that form between opposing strands of DNA. Ester bonds, which are flexible, bind the carbohydrate to the nucleobase. This bond is flexible and allows DNA strands to move and bend. The phosphate group is then joined to the sugar through phosphodiester bonds. Phosphodiester bonds are strong covalent bonds between one phosphate group and two carbon rings. These bonds act as the linkage between nucleotides to give the polymer defined structure and strength. Phosphodiester bonds are used to form the backbone in both DNA and RNA.
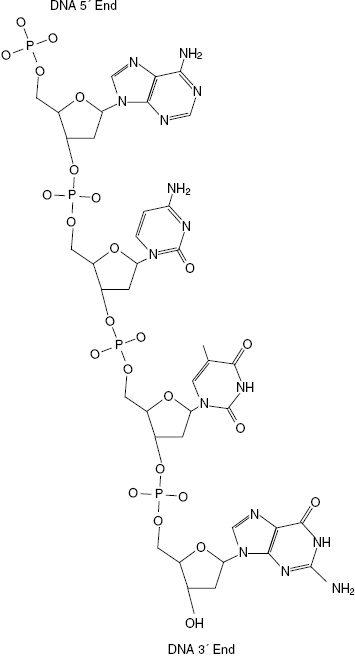
The backbone of DNA is a series of alternating carbohydrate and phosphate groups. Each phosphate of a nucleotide binds to the 5′ carbon of its sugar. The phosphate then binds to the next nucleotide at the 3′ carbon of the sugar. This rope-like structure is called a DNA polymer and, as mentioned earlier, is held together via phosphodiester bonds. Due to the asymmetric bonding nature of the phosphate groups, DNA and RNA molecules are given a direction. The direction is determined by the terminal end of the DNA strand. If a phosphate group is terminal, the phosphate is referred to as the 5′ end. However, if a hydroxyl group from the sugar is located at the DNA terminus, it is called the 3′ end. Shown in Fig. 6.5 is a picture of a single DNA polymer with labeled 5′ and 3′ ends. The directionality of the DNA polymer is important during transcription and replication, which is discussed later in this chapter.
Figure 6.5 Single DNA strand with 5′ and 3′ ends labeled. The 5′ end of DNA terminates with a phosphate group, see top of polymer, and the 3′ end terminates with a sugar group, see bottom of polymer.
As complementary DNA strands come together, they form a secondary structure that is similar to a ladder. The sides of the ladder, the DNA backbone, are alternating carbohydrate and phosphate groups. The polymers align, in opposing directions, with each other, and hydrogen bonds are formed between complementary nucleotides. As appropriate hydrogen bonds form, a right-handed double helix is formed. Each base pair, either G-C or A-T, appears as a rung on a ladder.
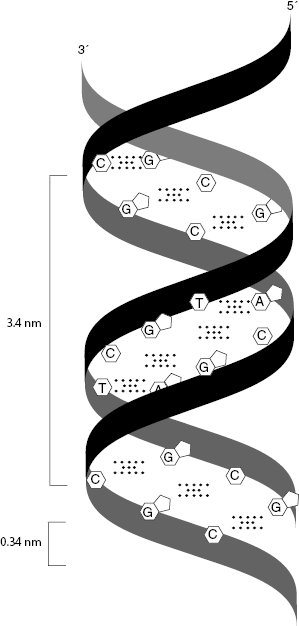
The DNA double helix includes two strands of DNA and the hydrogen bonds that connect complementary nucleobases. The sugar–phosphate backbone of each DNA polymer is on the outside of the helix, while the bases are on the inside. Each base pair is stacked 0.34 nm from the next and is added in a nearly perpendicular fashion to the long axis of the DNA polymer. As the helix is formed, each complete turn, approximately 10.5 base pairs, fills a 3.4-nm length. The spacing between each nucleotide pair and helical turn are shown in Fig. 6.6. This configuration is the regular structure of DNA found under normal physiological conditions, such as those within the nucleus. Additional DNA structures will not be discussed.
Figure 6.6 Spacing between each nucleotide pair, 0.34 nm, within a DNA helix, and the spacing, 3.4 nm, between complete helical turns are labeled.
Following organization of the nucleotides into DNA polymers and the double helix, DNA molecules still undergo further packaging. Chromosomal DNA molecules are much longer than the diameter of the nucleus itself and must be highly compacted. To begin the process, DNA is coiled around a series of histones. Histones are a set of eight proteins, known as an octomer. DNA winds around the surface of the histone in a helical pathway. The histone complex contains, on average, two complete turns of DNA, which consists of approximately 150 base pairs.
After DNA has successfully wrapped around the histone octamer, the DNA–histone complex is referred to as a nucleosome. Nucleosome structure is often compared with a string of beads. The string is free unwound DNA between the nucleosomes and the beads are DNA–histone complexes. A drawing of a nucleosome substructure is shown in Fig. 6.7. As can be seen, formation of nucleosomes reduces the accessibility of DNA to transcription and regulatory proteins. Both strands of DNA must be free for proper DNA binding. Protein factor binding of free DNA has been shown to be 10- to 104
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree