Fig. 1
Analysis of a natural seawater population. (a) Scatterplot of area vs. chlorophyll fluorescence as an example of a conventional cytometer output. Subpopulations are not readily distinguishable. (b) An aspect ratio vs. chlorophyll scatterplot, typical of imaging cytometry. Distinct subpopulations (R1 and R2) are visible on the plot, and examination of images identified additional subpopulations (R3–5). Colors of gates correspond to sets of images displayed at the periphery of the plot. Colored dots in (a) are the populations defined in (b), demonstrating the lack of resolution of area vs. chlorophyll
The ability to image allows documentation of variation between features of individual cells in a clonal population. Clonal populations under controlled culture conditions may be thought of as being homogeneous, however a variety of non-genetically based variations are possible [11–14]. Heterogeneity in clonal populations can be quantified by manifestation of an observable phenotype, either directly through morphological changes or through the use of dyes that serve as proxies for physiological state. The breadth of variation cannot be directly documented with bulk measurements.
Examples of heterogeneity in clonal populations demonstrate several important concepts. One example in microalgae is during accumulation of the neutral nonpolar lipid class triacylglycerol (TAG). TAG can be monitored by specific lipophilic dyes such as Nile Red [15, 16] and BODIPY [17]. In Fig. 2, we show the analysis of TAG accumulation after glucose addition to the culture medium in the chlorophytic algae Chlorella protothecoides, monitored by fluorescence of Nile Red. C. protothecoides reproduces by forming autospores, in which multiple daughter cells are contained within a single mother cell [18]. Imaging of cells enables a clear distinction between mother and daughter cells based on size and number of chloroplasts visible per cell (Fig. 2). We defined four categories (Fig. 2) based on cell type and lipid content and monitored TAG accumulation after 24 h of glucose treatment. Single daughter cells comprise 94 % of the population. A subpopulation of the daughter cells consisting of 20 % of the total cell number accumulated 36 % of the total TAG in the culture, but the remainder of the daughter cells (75 % of the total in the population) did not accumulate TAG (Fig. 2). Approximately 6 % of the total cell number was mother cells that accounted for 61 % of the total TAG accumulated in the culture, and a small fraction of the mother cells did not accumulate TAG (Fig. 2). This analysis highlights the ability of imaging cytometry to define different life stages of cells, particularly by enumerating the number of chloroplasts, and shows that the response of individual cells within a given life stage can respond dramatically differently from each other.
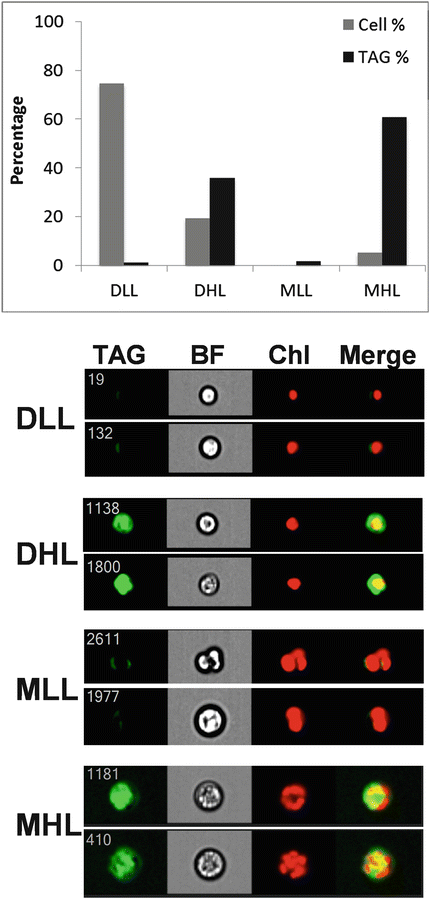

Fig. 2
Contribution of subpopulations of cells to TAG accumulation in Chlorella protothecoides. Four classes of cells were identified, Daughters Low Lipid (DLL), Daughters High Lipid (DHL), Mothers Low Lipid (MLL), and Mothers High Lipid (MHL), two examples of each class are shown at bottom. Images are TAG, bright field (BF), chlorophyll (Chl), and merged TAG and chlorophyll (Merge). Top, the percentage of cells in each class and their percentage contribution to total TAG in the culture are plotted
During the course of TAG accumulation, fluorescent staining can transition from diffuse (where TAG is dispersed within the cell), to concentrated (TAG-containing lipid droplets accumulate). Droplet sizes can also vary in distinct ways, from small droplets to large aggregates that constitute a majority of the cell volume. This is illustrated in Fig. 3, which monitors the accumulation of TAG in the diatom Cyclotella cryptica during a time course of silicon starvation [19]. Six distinct subpopulations were identified based on BODIPY and chlorophyll fluorescence (Fig. 3b). These different populations represent distinct phases in TAG accumulation and can be considered as phenotypes [19]. A useful feature of the IDEAS® software is that displayed images can be selected as those with the true average value of fluorescence of a particular population, as is shown in Fig. 3b. Population shifts based on cellular phenotypic differences can be used to make informed choices for other analyses. A bulk analysis of the experiment in Fig. 3 indicates a biphasic process of TAG accumulation with a slower phase between 0 and 48 h, followed by a much more rapid phase between 48 and 72 h (Fig. 3c). This suggests that the hyperaccumulating high chlorophyll subpopulation, which becomes dominant during this time (Fig. 3d), is undergoing a fundamentally different process of TAG accumulation than occurred with earlier subpopulations. Sampling time points between which substantial shifts in subpopulations occur would be highly informative in terms of understanding the mechanisms underlying the metabolic shifts. The additional phenotypic information provided by imaging makes this approach more informative than a bulk measurement or by conventional cytometry, where identifying the phenotype would be more cumbersome.
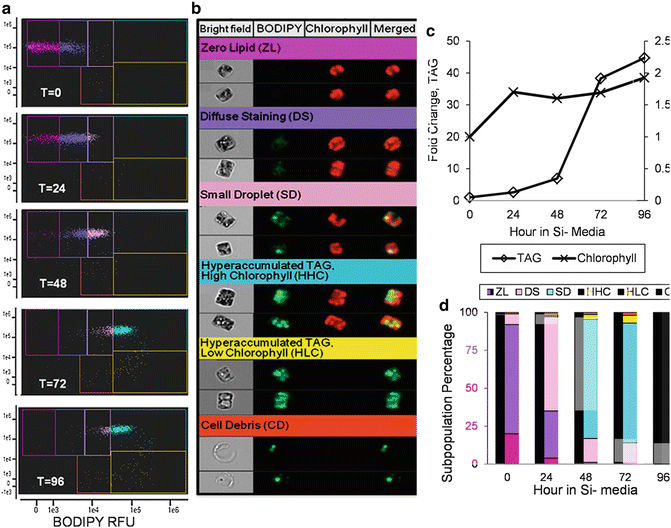

Fig. 3
Imaging cytometry analysis of TAG accumulation during silicon starvation of Cyclotella cryptica. A scatter plot time course analysis of changes in chlorophyll and BODIPY fluorescence (a) revealed six sub-populations of cells (b). (b) shows representative cells that are the actual mean for the subpopulation, and the color of the heading corresponds to a particular gated subpopulation in (a). Each cell has four images: bright field, BODIPY fluorescence (falsely colored green), chlorophyll autofluorescence (falsely colored red), and a merged image of BODIPY and chlorophyll. (c) Bulk measurement of the mean fold change in TAG and chlorophyll content per cell during the silicon starvation time course. (d) Relative percentage of cells within the different subpopulations during the time course. Colors of each bar correspond to the color of each gate in (a). Changes in the rate of TAG accumulation correspond to shifts into the HHC subpopulation. Figures are from Traller and Hildebrand [19]
Growth conditions or selection processes can generate subpopulations with overall morphology or size characteristics relevant to life cycle or growth processes; such information can be readily captured by imaging cytometry. Because diatoms have rigid silica-based cell walls that come in a wide variety of species-specific shapes, they are especially amenable to size and morphological analysis. With some diatom species that colonize surfaces, individual cells cannot be easily separated due to aggregation resulting from secretion of adhesive material. This prevents accurate morphological measurements of individual cells. Taking advantage of the silica cell wall, which can be cleaned free of organic material by acid treatment, we developed a method to image subpopulations of these cell types [20]. After acid cleaning [21] diatom silica can be fluorescently stained with APS-FITC [22, 23], and silica structures can subsequently be analyzed by imaging cytometry (Fig. 4). In Fig. 4, we identify two different sized subpopulations of the surface-associated diatom Craspedostauros australis by first identifying structures called valves in a scatterplot (Fig. 4a), and then plotting the valve length in a histogram (Fig. 4b). These clearly separate into large and small subpopulations (Fig. 4b–d).
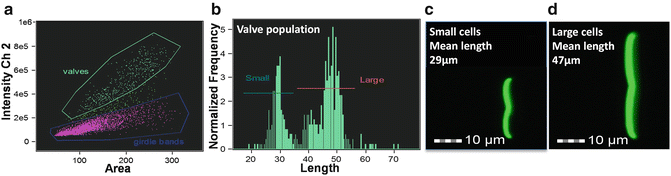

Fig. 4
Variation in C. australis cell wall morphology analyzed after acid-cleaning and staining with APS-FITC. (a) Scatterplot showing cell wall substructures called valves and girdle bands gated into separate populations. (b) Histogram of the length of the valves demonstrating two populations of different sizes. (c) Image of a representative cell from the bin with the mean value of the small population. (d) Image of a representative cell from the bin with the mean value of the large population
Another valuable application of imaging cytometry is in monitoring cellular processes by examining changes in cellular features. The location of intracellular components is an important aspect of understanding their function. Conventional cytometry lacks the ability to localize intracellular components, and conventional microscopy lacks high-throughput capability to generate robust statistics, as well as being subject to unintentional biases. In Fig. 5, we demonstrate the ability of IDEAS® software to distinguish the intracellular location of a GFP-tagged protein which, depending on conditions, is located in the chloroplast or in the endoplasmic reticulum (ER). A factor that complicates this analysis is that in diatoms the ER also surrounds the chloroplast [24]. The “co-localization” wizard in IDEAS® is able to distinguish between the two possibilities, most likely due to the distinct GFP fluorescence around the nucleus in the cells with ER localization (Fig. 5).
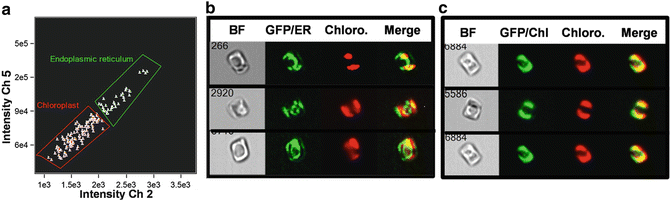

Fig. 5
Distinction of intracellular location of a GFP-tagged protein using the IDEAS software Co-localization wizard. (a) Scatterplot showing subpopulations identified as having endoplasmic reticulum (ER) or chloroplast localization. (b) Cell images with ER localization of the protein in bright field (BF), GFP, chloroplast (Chloro.), and merged channels. (c) Cell images with chloroplast localization of the protein with the same channels
An important cellular process is the partitioning of components during division. In Fig. 6, we document an unequal accumulation of TAG in dividing cells of C. cryptica, in which all visible lipid droplets are found in one daughter cell but not the other. In this analysis, a bright detail similarity feature for chlorophyll and BODIPY channels enables determination of the relative localization of probes within single cells. A histogram plot easily identifies those cells in which the bright detail of chlorophyll and BODIPY fluorescence are least similar (i.e., overlap the least).
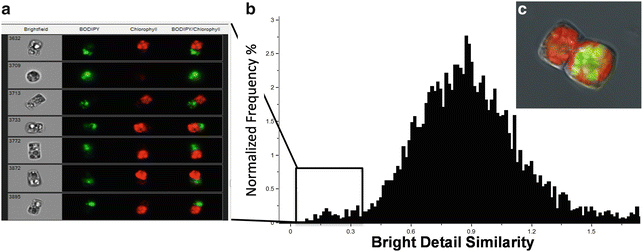

Fig. 6
Differential partitioning of chloroplasts and lipid droplets during cell division as shown by BODIPY and chlorophyll fluorescence. (a) Images of subpopulation of cells identified using the bright detail similarity feature of the two channels of interest to localize chlorophyll (red ) and lipid droplets (green) within one cell. Numbers in bright field denote the cell identifier. (c) High-resolution image taken using a fluorescence microscope showing this phenomenon
Another analysis of cellular components is monitoring organelle abundance and features. In Fig. 7, we applied imaging flow cytometry to characterize chloroplasts and their replication in diatoms. By defining a custom mask on chlorophyll fluorescence and utilizing the spot count feature in IDEAS®, a variety of aspects of chloroplasts can be characterized. These include the total number of chloroplasts per cell, the average area within a cell that is occupied by a chloroplast (a proxy for chloroplast size), and the fluorescence intensity per chloroplast (a proxy for the extent of chloroplast pigmentation). Using imaging flow cytometry during an experimental time course of silicon starvation in the diatom Thalassiosira pseudonana, chloroplast replication along with changes to chloroplast size and pigmentation could be documented (Fig. 7). We were able to quantify what fraction of the increase in bulk chlorophyll fluorescence was attributed to increases in the number, size, and pigmentation of chloroplasts.
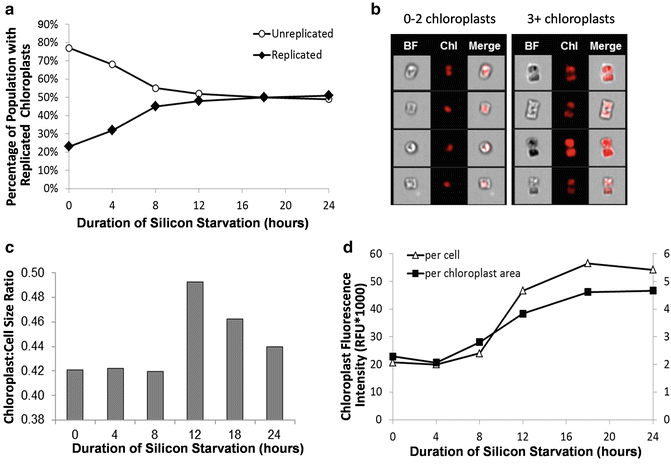

Fig. 7
Changes to chloroplast number, size, and fluorescence intensity in silicon-starved T. pseudonana determined by imaging flow cytometry. (a) Percentage of population with replicated chloroplasts. (b) Example images of cells with (n) chloroplasts. (c) Ratio of the average chloroplast size (μm2) to average cell size (μm2). (d) Average chloroplast fluorescence intensity normalized on a per cell basis and per chloroplast area
In some cases, the angle of imaging is important in analysis, especially with non-spherical cells. In Fig. 8, we show the analysis of silica incorporation into the cell walls of T. pseudonana, a cylindrically shaped diatom. Silica incorporation is monitored using the fluorescent dye PDMPO, which quantitatively incorporates into forming silica [25]. Different possible viewing angles of the cell are shown in Fig. 8a; the major features to consider are structures called valves and girdle bands. In this example, specific classifiers were developed to identify cells in different orientations. A custom classifier (the “Round” classifier) combined eight individual features in a proportional manner, as well as customized image masks and image algorithms using Boolean logic to distinguish diatoms in a valve vs. girdle band view (see Subheading 3). Even when cells are cultivated in the presence of excess dye, it is likely that a portion of the population will lack dye incorporation in both valves. These cells can only be detected when imaged in the girdle plane, and will have much lower mean fluorescence intensity than fully labeled cells. Cells imaged in valve view have slightly higher mean fluorescence intensity than cells imaged in girdle view (Fig. 8d). This is likely due to an additive effect of imaging fluorescence intensity of both valves and the girdle bands stacked on top of one another. The fluorescently labeled valves and girdle bands are viewed independently when cells are imaged in the girdle plane and display slightly lower mean fluorescence intensity (Fig. 8d).
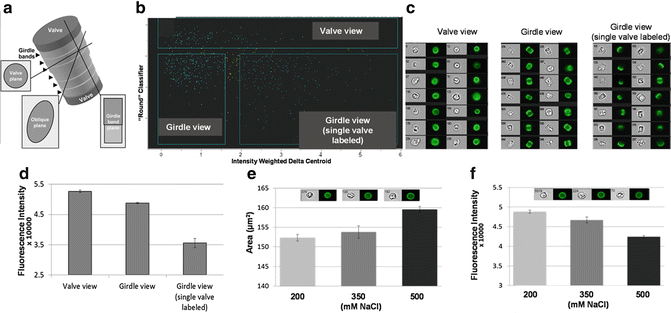

Fig. 8
Orientation effects on data interpretation. (a) Schematic of a cylindrically-shaped diatom cell, showing views at different angles. (b) Scatter plot of processed data to separate subpopulations of cells from different view angles. (c) Images of cells from each subpopulation in bright field and PDMPO fluorescence. (d) Fluorescence intensity of cells imaged in the three different views. (e) Area of valves from cells grown under three different NaCl concentrations. Inset shows representative cells from the three NaCl concentrations, indicating increasing valve diameter. (f) Fluorescence intensity of valves from cells grown under three different NaCl concentrations. Inset shows representative cells from the three NaCl concentrations, indicating decreasing PDMPO fluorescence
In Fig. 8e, we demonstrate slight, but statistically significant, changes in valve diameter during growth under different salinities. Interestingly, PDMPO fluorescence decreases with increasing salinity (Fig. 8f), indicating that valves are becoming either thinner or less dense under these conditions, which is consistent with previous findings based on other, more involved, methods [26, 27]. Without the robust statistics enabled by the custom classifier and large number of images, reaching this conclusion would not have been possible.
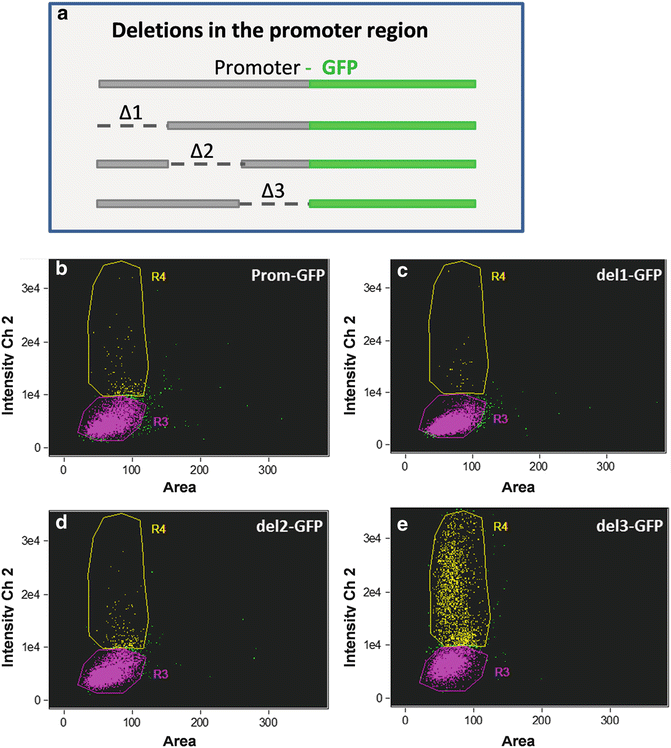
We have also applied ImageStream® analysis to the study of gene expression and epigenetic effects related to that. In Fig. 9, we document variation in cellular responses in a clonal population of a promoter deletion series by monitoring GFP expression. The data show a conserved subpopulation with similar GFP fluorescence in the intact control and with all three regions of deletion. In addition, in all constructs there is a subpopulation of cells with higher GFP fluorescence, but the percentage of cells in this subpopulation in del 3 is 12- to 85-fold higher than the others. This suggests the presence of a partially leaky repressor element in the del 3 region, which is functional only in a percentage of the population under the conditions tested. Substantial cell-to-cell variation in expression has been documented in other unicellular organisms or within the population of a particular cell type [11–13]. The ability to monitor levels of gene expression in individual cells within the population provides more accurate description regarding cellular response to various conditions than a bulk measurement. In Fig. 10, we monitored GFP fluorescence to evaluate the contribution of different 5′ and 3′ flanking regions to expression levels. The constructs included nitrate reductase (NR) and fucoxanthin chlorophyll binding protein (FCP) 5′ and 3′ flanking regions in various combinations (Fig. 10). NR was previously shown to respond to the presence or absence of nitrate, and to be repressed by ammonium [28], but it is unclear where the control elements mediating this response are located. In cells expressing GFP, the mean fluorescence was similar with the different constructs and conditions (Fig. 10a), indicating that the extent of the response was tightly constrained; that is, there was little tailoring of the level of response. The major difference was in the percentage of cells expressing GFP; the NR5′ flanking region was essential for expression in the majority of cells (Fig. 10b). However, even with NR 5′ and 3′ flanking regions only 50 % of the cells expressed GFP in the presence of nitrate, possibly suggesting that nitrogen status differs between cells. Control is exerted by both the 5′ and 3′ flanking region of NR, the presence of the 5′ flanking region enables expression in the majority of cells, but this is improved by replacing the 3′ flanking region with that of FCP, suggesting that a repression element may be present in this portion of the NR 3′ flanking region or a stimulating element is present in the 3′ flanking region of FCP.